The cassandra paradox: looking into the crystal Ball of radiomics in thoracic surgery
INTRODUCTION
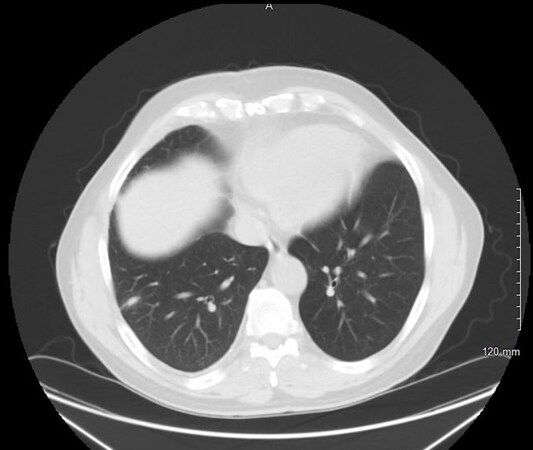
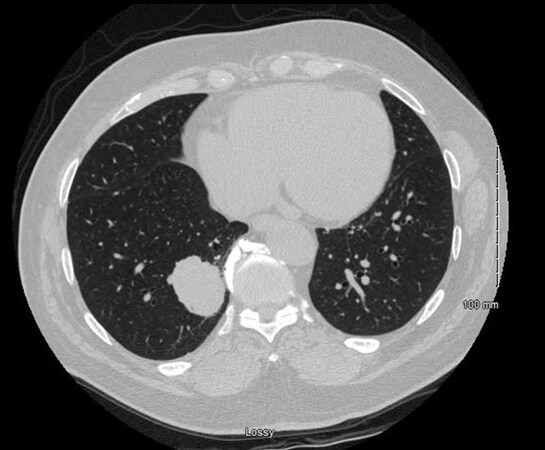
Lung cancer is the leading cause of cancer-associated death in both men and women. Research is being conducted to identify best practices for screening and diagnosing this morbid disease, and we continue to rely on computed tomography (CT) and positron emission tomography (PET)/CT scans as our modalities of choice. Historically, nodule size has been the single-most important determinant of therapeutic options as lung cancer increases exponentially with nodule size[1-6]. Radiomics is a rapidly progressing field of study that extracts quantitative variables and identifies patterns that are difficult to evaluate on the routine reading of CT or PET/CT images. Some studies have shown radiomics parameters to correlate with tumor aggressiveness[7] as well as a prognostic determinant of survival[8-11]. Given the benefits of this potential technology, clinicians hope to derive quantitative features from these images that can make more accurate diagnoses, guide patient-specific treatment options, and potentially predict tumor behavior with respect to treatment response and survival[12]. As an example, radiomics may help distinguish that the lung nodule in Figure 1 is a malignant adenocarcinoma of the lung with more metastatic potential than the larger lung nodule seen in Figure 2, which is a less aggressive typical carcinoid. Clinicians using this technology could become modern day Cassandras. Greek mythology explains that Cassandra was gifted with the knowledge of future events. Radiomics offers the ability for fortune telling through a modern crystal ball of radiology and computational informatics.
CT images are digital images that consist of pixels when viewed in 2 dimensions and voxels when viewed in 3 dimensions. Radiomic features are derived from the pixels and can be subdivided into four distinct groups (statistical, model-based, transform-based, and shape-based)[13]. Statistical based features such as grey-level mean, max, min, variance, and percentiles are based on a single pixel, or voxel analysis. Gray-level cooccurrence matrix (GLCM), gray-level size zone matrix, gray-level run-length matrix (NGTDM), and neighborhood gray-level dependence matrix evaluate the spatial relationship of multiple voxels[7]. Fractal analysis, a model-based feature, illustrates the structural detail of a lesion with improved resolution. A transform-based feature is wavelet decomposition, which involves passing the image through multiple filters[14]. Many of these parameters describe the “texture” of the region of interest. In essence, they are describing the changes in the grayscale (neighboring pixels) within the region being evaluated. Deriving radiomic parameters is beyond the scope of our analysis, and we refer the reader to the references cited. We aim to evaluate the feasibility, identify possible applications, and project a future for radiomics in the field of thoracic surgery.
RADIOMICS ANALYSIS
Pathologic diagnoses have previously been established with tissue biopsies. Peripheral endobronchial ultrasound with a fine needle aspiration is frequently used as a diagnostic modality for central nodules that can be identified. However, Gerlinger et al.[15] showed that random samples of tumor tissues acquired through this method may fail to accurately represent the breadth of biological variation within tumors. Radiomics are able to extract a substantially greater number of nodule features with much better reproducibility compared to CT and PET/CT scans[16]. Radiomics attempts to identify quantitative imaging characteristics from images in order to non-invasively predict the behavior of tumors[17,18]. This genomic heterogeneity is inadequately evaluated via CT or PET/CT because of the partial volume effect[19,20]. The heterogeneity of a tumor as a result of variations in gene expression is a common feature of malignant tumors[21-24]. Petkovska et al.[25] in 2006 showed that GLCM extracted from contrast-enhanced CT scans can accurately identify malignant from benign nodules more accurately than three experienced radiologists. A nodule’s volume doubling time can estimate the likelihood of malignancy as indicated in the NELSON trial[26]. This method requires proper surveillance and a longer amount of time prior to official diagnosis than the use of radiomics. Hawkins et al.[27] identified 23 stable features that could predict nodules that would become cancerous 1 and 2 years hence with accuracies of 80% and 79%.
When Lee et al.[28] 2014 used texture analysis in combination with clinical and CT features; differentiating power between benign and malignant lesions increased significantly compared to clinical and CT features alone. Analysis of these nodules has the potential to identify pathognomonic features of malignant disease. Kido et al.[29] 2002 showed that the fractal dimensions for bronchogenic carcinomas were significantly smaller than pneumonias and tuberculomas (P < 0.0001). Other studies were able to identify radiomic features specific for EGFR mutant vs. wild type groups and K-ras mutations[30-32]. CANARY, an application designed to stratify lung adenocarcinoma into aggressive and minimally invasive, uses nine representative characteristics to identify the patient’s histopathology[33]. Dong et al.[34] 2013 found standardized uptake value (SUV), maximum standardized uptake value (SUV MAX), GLCM-entropy, and GLCM-energy were found to be significantly correlated with T and N stage.
Predictive models that combined radiomic features significantly improved the prediction of pathologic response to therapy, according to Yang et al.[35] 2014. This was further supported by Cook et al.[36] in 2013, who found that neighborhood gray tone difference matrix (NGTDM) derived coarseness, busyness, and contrast could predict response to chemoradiotherapy better than SUVs as reported by a PET/CT scan. Additionally, Coroller et al.[37] in 2016 identified seven radiomic features that were predictive for pathologic gross residual disease and one for pathologic complete response. Wang et al.[38] identified eight features using LASSO Cox analysis to develop a radiomic signature used to predict recurrence-free survival. They found that histology (P < 0.001) and their radiomics signature (P < 0.001) were independent prognostic factors for recurrence free survival. The advantages of radiomics are clear as it would obviate the need for biopsies, which are invasive and often unsuccessful. Using radiomics to predict genomic data (radiogenomics) would allow better profiling of tumors as sampling errors would potentially be avoided as well as for the need for re-biopsy as tumors change in response to therapy over time.
RADIOMICS IN THE FUTURE
Although there is considerable promise in the field of radiomics, there are clear challenges in the burgeoning field. There is considerable variation in the conduct of radiologic examinations. Variation can occur not only among different institutions and machines but also on the same machine on successive scans[16]. Radiologic exam variation can make interpretation of the data challenging generating too much noise to differentiate any meaningful clinical parameters. Terminology for various radiomic metrics is also incompletely standardized, leading to inconsistencies in data collection and interpretation. Many radiomic features are redundant, which can lead to both overfitting and underfitting[7]. Overfitting occurs when a model with a large dataset and a high degree of freedom remembers all data points, thus making it more difficult to differentiate disease-specific features from noise; underfitting creates an overly simplistic model. Published radiomic studies often have small sample sizes that lack a separate validation dataset to test the constructed model. Clinical relevance of radiomic models is often lacking with poor collaboration between clinicians and modelers.
As radiomics continues to evolve and become more widely accepted, additional prospective studies will be required to evaluate its feasibility. Large trials such as the National Lung Cancer Screening Trial (NLST) and lung cancer databases such as Qualitative Imaging Network (QIN), National Cancer Institute (NCI), and others could be mined to uncover possible associations between imaging characteristics and tumor histopathology. Standard acquisition parameters and image processing methods should be adopted; some studies recommend small voxels, narrow gaussian postfiltering, and point-spread function modeling[39,40]. Additionally, inter-observer variability will need to be optimized as this is a common variation within the field of radiation oncology. Pavic et al.[41] evaluated the impact of inter-observer variability on the accuracy and generalizability of radiomic results. They found that manual tumor delineation was variable and observer-dependent, thus leading to a reduction in the number of applicable radiologic features. Deep learning networks avoid the need to manipulate images to identify radiomic criteria personally. These neural networks are taught to identify and discriminate between objective data points within the image. Using deep learning, da Silva et al.[42] were able to identify small lung nodules with an accuracy of 97.6%. In combination with radiomics, features identified by a pre-trained deep learning network were 90% accurate in the prediction of survival of patients with adenocarcinoma[43]. Avanzo et al.[44] postulate that deep learning will “facilitate faster clinical translation of lung cancer radiomics” given the unmet clinical need for lung cancer diagnosis and surveillance.
Due to the increased surveillance of lung nodules and trials such as the NLST, the use of segmentectomy over lobectomy for early-stage non-small cell lung cancer has become more widely accepted[45-53]. Segmentectomies are notoriously difficult because of the lack of anatomic differentiation of bronchioles and vasculature. A pilot study out of the Netherlands showed the feasibility of a virtual reality-based application that uses artificial intelligence to preoperatively plan a segmentectomy[54]. Four of the ten patients evaluated in this study were found to have final pathology that was discordant with preoperative diagnosis, which could be clinically significant. One patient’s final pathology was benign, while the preoperative diagnosis was suspicious for metastasis. Intraoperatively, through the use of indocyanine green, intersegmental borders can be identified, as shown by Iizuka et al.[55]. These platforms could change the way that patients are preoperatively evaluated and provide a more patient-specific resection.
CONCLUSION
Radiomics is a developing field that has the potential to radically change the diagnosis and management of lung nodules and lung cancer. However, the power of Cassandra comes with a potential problem. Cassandra was cursed by Apollo, who prevented others from believing her prophecy. Will clinicians and patients believe the diagnoses and treatment algorithms predicted by radiomics? Or will they still require a tissue diagnosis? Will the predictive power of radiomics lead to other unintended consequences known or unknown? Regardless, advances in imaging technology, computational power, and artificial intelligence will hopefully drive the field forward to create a powerful clinical useful modality.
DECLARATIONS
Authors’ contributionsMade major contributions to the conception, drafting, and editing of the manuscript: Decker JM
Conception and editing of the manuscript: Sesti J
Made major contributions to the editing and formating of the manuscript: Turner AL
Supervised the conception, writing, and editing of the manuscript: Paul S
Availability of data and materialsNot applicable.
Financial support and sponsorshipNone.
Conflicts of interestAll authors declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2022.
REFERENCES
1. Ko JP, Berman EJ, Kaur M, et al. Pulmonary Nodules: growth rate assessment in patients by using serial CT and three-dimensional volumetry. Radiology 2012;262:662-71.
2. Henschke CI, Yankelevitz DF, Yip R, et al. Writing Committee for the I-ELCAP Investigators. Lung cancers diagnosed at annual CT screening: volume doubling times. Radiology 2012;263:578-83.
3. Hasegawa M, Sone S, Takashima S, et al. Growth rate of small lung cancers detected on mass CT screening. Br J Radiol 2000;73:1252-9.
4. Wilson DO, Ryan A, Fuhrman C, et al. Doubling times and CT screen-detected lung cancers in the Pittsburgh Lung Screening Study. Am J Respir Crit Care Med 2012;185:85-9.
5. Jennings SG, Winer-Muram HT, Tann M, Ying J, Dowdeswell I. Distribution of stage I lung cancer growth rates determined with serial volumetric CT measurements. Radiology 2006;241:554-63.
6. Winer-Muram HT, Jennings SG, Tarver RD, et al. Volumetric growth rate of stage I lung cancer prior to treatment: serial CT scanning. Radiology 2002;223:798-805.
7. Sala E, Mema E, Himoto Y, et al. Unravelling tumour heterogeneity using next-generation imaging: radiomics, radiogenomics, and habitat imaging. Clin Radiol 2017;72:3-10.
8. Yang F, Wang Y, Li Q, et al. Intratumor heterogeneity predicts metastasis of triple-negative breast cancer. Carcinogenesis 2017;38:900-9.
9. Burrell RA, McGranahan N, Bartek J, Swanton C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 2013;501:338-45.
10. Liu J, Dang H, Wang XW. The significance of intertumor and intratumor heterogeneity in liver cancer. Exp Mol Med 2018;50:e416.
11. Morris LG, Riaz N, Desrichard A, et al. Pan-cancer analysis of intratumor heterogeneity as a prognostic determinant of survival. Oncotarget 2016;7:10051-63.
12. Mayerhoefer ME, Materka A, Langs G, et al. Introduction to radiomics. J Nucl Med 2020;61:488-95.
13. Benoit-Cattin H. Texture analysis for magnetic resonance imaging. Available from: https://books.google.com.hk/books?hl=zh-CN&lr=&id=kNFKek4RtnkC&oi=fnd&pg=PA9&dq=Texture+Analysis+for+Magnetic+Resonance+Imaging&ots=PGtzjqs0Sy&sig=cQPfAbJy7KIqWCXJEijzRLabik4&redir_esc=y#v=onepage&q=Texture%20Analysis%20for%20Magnetic%20Resonance%20Imaging&f=false [Last accessed on 21 Mar 2022].
14. Laine A, Fan J. Texture classification by wavelet packet signatures. IEEE Trans Pattern Anal Machine Intell 1993;15:1186-91.
15. Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 2012;366:883-92.
16. Wilson R, Devaraj A. Radiomics of pulmonary nodules and lung cancer. Transl Lung Cancer Res 2017;6:86-91.
17. Gillies RJ, Kinahan PE, Hricak H. Radiomics: images are more than pictures, they are data. Radiology 2016;278:563-77.
18. Lambin P, Rios-Velazquez E, Leijenaar R, et al. Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer 2012;48:441-6.
19. Soret M, Bacharach SL, Buvat I. Partial-volume effect in PET tumor imaging. J Nucl Med 2007;48:932-45.
20. Hatt M, Cheze-le Rest C, van Baardwijk A, Lambin P, Pradier O, Visvikis D. Impact of tumor size and tracer uptake heterogeneity in (18)F-FDG PET and CT non-small cell lung cancer tumor delineation. J Nucl Med 2011;52:1690-7.
21. Maley CC, Galipeau PC, Finley JC, et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat Genet 2006;38:468-73.
22. Marusyk A, Almendro V, Polyak K. Intra-tumour heterogeneity: a looking glass for cancer? Nat Rev Cancer 2012;12:323-34.
23. Chicklore S, Goh V, Siddique M, Roy A, Marsden PK, Cook GJ. Quantifying tumour heterogeneity in 18F-FDG PET/CT imaging by texture analysis. Eur J Nucl Med Mol Imaging 2013;40:133-40.
24. Fisher R, Pusztai L, Swanton C. Cancer heterogeneity: implications for targeted therapeutics. Br J Cancer 2013;108:479-85.
25. Petkovska I, Shah SK, McNitt-Gray MF, et al. Pulmonary nodule characterization: a comparison of conventional with quantitative and visual semi-quantitative analyses using contrast enhancement maps. Eur J Radiol 2006;59:244-52.
26. Horeweg N, van der Aalst CM, Vliegenthart R, et al. Volumetric computed tomography screening for lung cancer: three rounds of the NELSON trial. Eur Respir J 2013;42:1659-67.
27. Hawkins S, Wang H, Liu Y, et al. Predicting malignant nodules from screening CT scans. J Thorac Oncol 2016;11:2120-8.
28. Lee SH, Lee SM, Goo JM, Kim KG, Kim YJ, Park CM. Usefulness of texture analysis in differentiating transient from persistent part-solid nodules(PSNs): a retrospective study. PLoS One 2014;9:e85167.
29. Kido S, Kuriyama K, Higashiyama M, Kasugai T, Kuroda C. Fractal analysis of small peripheral pulmonary nodules in thin-section CT: evaluation of the lung-nodule interfaces. J Comput Assist Tomogr 2002;26:573-8.
30. Liu Y, Kim J, Balagurunathan Y, et al. Radiomic features are associated with EGFR mutation status in lung adenocarcinomas. Clin Lung Cancer 2016;17:441-8.e6.
31. Weiss GJ, Ganeshan B, Miles KA, et al. Noninvasive image texture analysis differentiates K-ras mutation from pan-wildtype NSCLC and is prognostic. PLoS One 2014;9:e100244.
32. Yip SS, Kim J, Coroller TP, et al. Associations between somatic mutations and metabolic imaging phenotypes in non-small cell lung cancer. J Nucl Med 2017;58:569-76.
33. Maldonado F, Boland JM, Raghunath S, et al. Noninvasive characterization of the histopathologic features of pulmonary nodules of the lung adenocarcinoma spectrum using computer-aided nodule assessment and risk yield (CANARY)--a pilot study. J Thorac Oncol 2013;8:452-60.
34. Dong X, Xing L, Wu P, et al. Three-dimensional positron emission tomography image texture analysis of esophageal squamous cell carcinoma: relationship between tumor 18F-fluorodeoxyglucose uptake heterogeneity, maximum standardized uptake value, and tumor stage. Nucl Med Commun 2013;34:40-6.
35. Yang F, Thomas MA, Dehdashti F, Grigsby PW. Temporal analysis of intratumoral metabolic heterogeneity characterized by textural features in cervical cancer. Eur J Nucl Med Mol Imaging 2013;40:716-27.
36. Cook GJ, Yip C, Siddique M, et al. Are pretreatment 18F-FDG PET tumor textural features in non-small cell lung cancer associated with response and survival after chemoradiotherapy? J Nucl Med 2013;54:19-26.
37. Coroller TP, Agrawal V, Narayan V, et al. Radiomic phenotype features predict pathological response in non-small cell lung cancer. Radiother Oncol 2016;119:480-6.
38. Wang T, Deng J, She Y, et al. Radiomics signature predicts the recurrence-free survival in stage I non-small cell lung cancer. Ann Thorac Surg 2020;109:1741-9.
39. Papp L, Rausch I, Grahovac M, Hacker M, Beyer T. Optimized feature extraction for radiomics analysis of 18F-FDG PET imaging. J Nucl Med 2019;60:864-72.
40. Lasnon C, Majdoub M, Lavigne B, et al. 18F-FDG PET/CT heterogeneity quantification through textural features in the era of harmonisation programs: a focus on lung cancer. Eur J Nucl Med Mol Imaging 2016;43:2324-35.
41. Pavic M, Bogowicz M, Würms X, et al. Influence of inter-observer delineation variability on radiomics stability in different tumor sites. Acta Oncol 2018;57:1070-4.
42. da Silva GLF, Valente TLA, Silva AC, de Paiva AC, Gattass M. Convolutional neural network-based PSO for lung nodule false positive reduction on CT images. Comput Methods Programs Biomed 2018;162:109-18.
43. Paul R, Hawkins SH, Balagurunathan Y, et al. Deep feature transfer learning in combination with traditional features predicts survival among patients with lung adenocarcinoma. Tomography 2016;2:388-95.
44. Avanzo M, Stancanello J, Pirrone G, Sartor G. Radiomics and deep learning in lung cancer. Strahlenther Onkol 2020;196:879-87.
45. Koike T, Yamato Y, Yoshiya K, Shimoyama T, Suzuki R. Intentional limited pulmonary resection for peripheral T1 N0 M0 small-sized lung cancer. J Thorac Cardiovasc Surg 2003;125:924-8.
46. Keenan RJ, Landreneau RJ, Maley RH Jr, et al. Segmental resection spares pulmonary function in patients with stage I lung cancer. Ann Thorac Surg 2004;78:228-33; discussion 228-33.
47. Harada H, Okada M, Sakamoto T, Matsuoka H, Tsubota N. Functional advantage after radical segmentectomy versus lobectomy for lung cancer. Ann Thorac Surg 2005;80:2041-5.
48. Okada M, Nishio W, Sakamoto T, et al. Effect of tumor size on prognosis in patients with non-small cell lung cancer: the role of segmentectomy as a type of lesser resection. J Thorac Cardiovasc Surg 2005;129:87-93.
49. Yoshida J, Nagai K, Yokose T, et al. Limited resection trial for pulmonary ground-glass opacity nodules: fifty-case experience. J Thorac Cardiovasc Surg 2005;129:991-6.
50. Altorki NK, Yip R, Hanaoka T, et al. I-ELCAP Investigators. Sublobar resection is equivalent to lobectomy for clinical stage 1A lung cancer in solid nodules. J Thorac Cardiovasc Surg 2014;147:754-62; Discussion 762-4.
51. Dziedzic R, Zurek W, Marjanski T, et al. Stage I non-small-cell lung cancer: long-term results of lobectomy versus sublobar resection from the Polish National Lung Cancer Registry. Eur J Cardiothorac Surg 2017;52:363-9.
52. Kates M, Swanson S, Wisnivesky JP. Survival following lobectomy and limited resection for the treatment of stage I non-small cell lung cancer<=1 cm in size: a review of SEER data. Chest 2011;139:491-6.
53. Wisnivesky JP, Henschke CI, Swanson S, et al. Limited resection for the treatment of patients with stage IA lung cancer. Ann Surg 2010;251:550-4.
54. Sadeghi AH, Maat APWM, Taverne YJHJ, et al. Virtual reality and artificial intelligence for 3-dimensional planning of lung segmentectomies. JTCVS Tech 2021;7:309-21.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Decker JM, Sesti J, Turner AL, Paul S. The cassandra paradox: looking into the crystal Ball of radiomics in thoracic surgery. Art Int Surg 2022;2:57-63. http://dx.doi.org/10.20517/ais.2022.05
AMA Style
Decker JM, Sesti J, Turner AL, Paul S. The cassandra paradox: looking into the crystal Ball of radiomics in thoracic surgery. Artificial Intelligence Surgery. 2022; 2(2): 57-63. http://dx.doi.org/10.20517/ais.2022.05
Chicago/Turabian Style
Decker, Jonathan M., Joanna Sesti, Amber L. Turner, Subroto Paul. 2022. "The cassandra paradox: looking into the crystal Ball of radiomics in thoracic surgery" Artificial Intelligence Surgery. 2, no.2: 57-63. http://dx.doi.org/10.20517/ais.2022.05
ACS Style
Decker, JM.; Sesti J.; Turner AL.; Paul S. The cassandra paradox: looking into the crystal Ball of radiomics in thoracic surgery. Art. Int. Surg. 2022, 2, 57-63. http://dx.doi.org/10.20517/ais.2022.05
About This Article
Copyright
Data & Comments
Data
 Cite This Article 6 clicks
Cite This Article 6 clicks











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.