Artificial intelligence for decision support in surgical oncology - a systematic review
Abstract
Aim: We systematically review current clinical applications of artificial intelligence (AI) that use machine learning (ML) methods for decision support in surgical oncology with an emphasis on clinical translation.
Methods: MEDLINE, Web of Science, and CENTRAL were searched on 19 January 2021 for a combination of AI and ML-related terms, decision support, and surgical procedures for abdominal malignancies. Data extraction included study characteristics, description of algorithms and their respective purpose, and description of key steps for scientific validation and clinical translation.
Results: Out of 8302 articles, 107 studies were included for full-text analysis. Most of the studies were conducted in a retrospective setting (n = 105, 98%), with 45 studies (42%) using data from multiple centers. The most common tumor entities were colorectal cancer (n = 35, 33%), liver cancer (n = 21, 20%), and gastric cancer (n = 17, 16%). The most common prediction task was survival (n = 36, 34%), with artificial neural networks being the most common class of ML algorithms (n = 52, 49%). Key reporting and validation steps included, among others, a complete listing of patient features (n = 95, 89%), training of multiple algorithms (n = 73, 68%), external validation (n = 13, 12%), prospective validation (n = 2, 2%), robustness in terms of cross-validation or resampling (n = 89, 83%), treatment recommendations by ML algorithms (n = 9, 8%), and development of an interface (n = 12, 11%).
Conclusion: ML for decision support in surgical oncology is receiving increasing attention with promising results, but robust and prospective clinical validation is mostly lacking. Furthermore, the integration of ML into AI applications is necessary to foster clinical translation.
Keywords
INTRODUCTION
Cancer is still a major problem in modern medicine[1], with surgery being an important part of curative multimodal treatment strategies for solid cancers[2]. Furthermore, operations on abdominal organs can be associated with many severe complications[3]. Choices regarding the optimal treatment for individual patients are made by multidisciplinary tumor boards that have to follow international guidelines[4-6], but even this multidisciplinary approach does not always guarantee treatment success[7].
Attempts to improve curative surgical treatment increasingly involve the use of modern computational methods[8]. Digitalization, interconnectivity between technical equipment, and electronic health records offer chances to improve patient outcomes. With the introduction of machine learning (ML) and artificial intelligence (AI) into medicine, multiple new options for data analysis have arisen and could be used to facilitate decisions in surgical oncology[9]. AI is intelligence demonstrated by machines, which may be realized by the use of ML, i.e., the study of computer algorithms that can improve automatically through experience[10]. While a general AI that can transfer knowledge to other problems similar to human intelligence is not yet realized, ML algorithms can be used to solve specific problems[11], such as interpretation of electrocardiograms[12], detection of suspicious findings in chest radiographs[13], computer-assisted colonoscopy[14], and early detection of pancreatic cancer[15].
Regarding a systematic analysis of available evidence in AI and ML for decision support, a recent systematic review investigated ML and regression analysis from an epidemiologist’s point of view, focusing on the algorithm performance and not on the clinical translation[16]. Another review assessed studies that used AI for analysis, detection, prediction, and pathology in gastric cancer[17]. A scoping review described the use of ML in metabolic surgery[18]. Other reviews focused on the application of data science methods to surgery[9] and general hurdles for clinical translation[19], or the use of ML in abdominal surgery[20] and surgical phase recognition[21].
In contrast, this systematic review gives a detailed overview of AI for decision support in surgical oncology based on ML with a focus on clinical applications, methodological soundness, and performance evaluation, as well as the key steps for clinical translation.
METHODS
The systematic review was conducted according to the preferred reporting items for systematic reviews and meta-analysis (PRISMA, see checklist in Supplementary Table 1)[22] and followed the recommendations of the Cochrane Handbook for Systematic Reviews and Interventions[23] and recommendations specific to systematic reviews in surgery[24]. A protocol for this systematic review was developed a priori and was registered in the PROSPERO database (CRD42021235515).
Eligibility criteria
Studies were selected following the criteria for population, intervention, control, and outcome (PICO criteria) described below. Animal studies, meeting abstracts, letters, comments, editorials, non-English literature, and publications for which the full text was irretrievable were excluded.
For the population, inclusion criteria were to meet the conditions of all studies on surgical oncology, including patients with the following types of cancer: thyroid, esophagus, stomach, colorectal, gallbladder, liver, pancreas, kidney, spleen, or sarcomas. Patients were eligible for, received, or followed up after cancer surgery. Excluded were other cancer types or a treatment approach that did not involve surgery.
For intervention, inclusion criteria were development, testing, and use of AI based on ML only for decision support to aid in diagnosis, prediction of tumor characteristics and prognosis, therapy planning, intraoperative, and postoperative problems. Excluded were decision support applications using imaging or biomarkers only without taking clinical characteristics into account, ML algorithms used for abdominal cancer screening only, and algorithm development without clinical application.
For control, inclusion criteria were based on comparison with other ML algorithms and regression analysis, if described. These were not mandatory for inclusion without specific exclusion criteria.
For outcome, inclusion criteria were descriptive characteristics of selected studies and analysis of quality for developing meaningful clinical decision support without specific exclusion criteria.
Definition of machine learning and artificial intelligence
There is no consensus on the definition of ML and AI, with some authors using the terms synonymously[25]. In this review, ML is defined “as a program that learns to perform a task or make a decision automatically from data”, according to Beam and Kohane[26]. Algorithms range from rule-based systems and regression analysis, which require more human input, to higher-ranking systems such as random forests or neural networks, which require less[26]. As ML has gained more popularity in medicine in recent years, the focus is on novel methods for decision support and excluded studies, which dealt with lower ranking regression analysis from the current review.
Poole et al. used the term “computational intelligence” as a synonym for artificial intelligence (AI), which is intelligence demonstrated by machines that perceive their environment, adapt their actions accordingly, and learn from experience[27]. Based on this definition, we focus on learning systems, and, for the purpose of this review, we define AI in surgical oncology as a computer application that integrates ML into the clinical workflow to support surgical decision making. These applications integrate ML algorithms trained on clinical data and surgical experience in a graphical user interface that allows for quick capture and representation of data in the clinical routine or even provides a treatment recommendation.
Information sources and search
The search strategy involved the most relevant medical databases for surgical literature; MEDLINE (via PubMed), Web of Science, and Cochrane Central Register of Controlled Trials (CENTRAL)[24]. The exact search terms can be accessed in Supplementary Note 1. The search was performed on 31 January 2021 for papers published between 1 January 2011 and 31 December 2020, because previously published applications of ML methods would not have been compared to current methods and would be outdated.
Study selection
After the removal of duplicates, titles and abstracts were screened by one reviewer (Haselbeck-Köbler M) for relevance. Positively evaluated reports were further screened by Schulze A, who provided an independent decision on whether to perform full-text screening. Selected articles underwent a full-text screening by two independent reviewers (Haselbeck-Köbler M and Schulze A). Any disagreement among the reviewers was resolved by consensus or a third person (Wagner M).
Data collection process
The data were extracted using a predefined extraction form which was refined after a pilot phase of data extraction for the first ten selected articles. Data extraction was performed by Haselbeck-Köbler M and Schulze A. Uncertainties were resolved by consensus or consultation of an independent third person (Wagner M).
Data were extracted for the bibliographic information of the selected articles, including title, year of publication, and first author. Furthermore, tumor entities were extracted, as well as the prediction task for which decision support was developed.
For the data used to construct ML algorithms, data origin (e.g., own clinical database or public registry database) was extracted, as well as the involvement of different facilities (single center or multiple centers), which categories of data were used (demographic, clinical, surgical data, laboratory, radiology, pathology, radiomics, and biomarker), and whether all variables used for machine learning were described. Studies were only defined as prospective studies if a fully developed ML algorithm for decision support was validated on new, incoming patients. Consequently, studies were defined as retrospective if the ML algorithms used prospectively collected data for training but were trained retrospectively.
For the decision support, the developed algorithms were extracted, and for the best algorithm, the performance metric “area under the receiver operating characteristic curve” (AUC) was extracted. For the development process of the algorithms, the total number of patients and, as surrogate measures for robustness, number of patients for external validation, process for data splitting or resampling, handling of missing values, and whether class imbalance was mentioned and adjusted were extracted. In addition, whether the code was available as open source was extracted.
For clinical translation, we extracted whether any kind of interface was developed and whether a demo was made accessible online to get an impression of the usability. In addition, whether a treatment recommendation was given instead of only a prediction that would not influence clinical decisions was extracted. A short summary of each paper was given with further specification of the clinical application.
Data were extracted on whether the criterion was met. Due to study heterogeneity, quantitative data synthesis was not viable because multiple cancer types and different algorithm purposes were evaluated, which would not allow comparisons with each other. Additionally, no risk-of-bias assessment was conducted, because the aim of this review was not to give clinical recommendations but to outline the research field.
Role of the funding source
This work was funded by the National Center for Tumor Diseases (NCT) Heidelberg, Germany, within the cancer therapy program “Surgical Oncology”, the German Federal Ministry of Health within the “Surgomics” project (grant number BMG2520DAT82), and the German Research Foundation (DFG, Deutsche Forschungsgemeinschaft) as part of Germany’s Excellence Strategy [EXC 2050/1, Project ID 3900696704 - Cluster of Excellence “Centre for Tactile Internet with Human-in-the-Loop” (CeTI)]. The funding sources did not influence the study design, collection, analysis, and interpretation of data; writing of the report; or the decision to submit the paper for publication.
RESULTS
The results of this systematic review include information about combining the expertise from surgery, machine learning, and interaction design to create a clinically usable AI for decision support in surgical oncology [Figure 1]. Accordingly, the extracted information was aggregated into a checklist of key steps for scientific validation and clinical translation of AI for decision support in surgical oncology based on the CHARMS Checklist for a systematic review of prediction models[28], an epidemiological literature review on ML prediction models[16], and an extensive discussion among the authors. Table 1 gives an overview of the steps together with their fulfillment in the studies included in this review, examples of fulfilling studies, and the necessary expertise from surgery, machine learning, and interaction design.
Figure 1. Necessary components for clinically usable artificial intelligence applications (AI). To create a clinically usable artificial intelligence application (AI) for decision support in surgical oncology, it is necessary to combine expertise from surgery, machine learning, and interaction design.
Checklist of key steps for scientific validation and clinical translation
| Step | Description | Fulfilled in studies | Examples | Necessary expertise [Figure 1] |
| Full feature set | Describe all patient characteristics used for building the algorithm, even before the selection of features for ML. For optimal comprehensibility, state whether the features were categorical or continuous, how they were transformed and describe at which point in time they were assessed and how | 89 % (n = 95) | Stojadinovic et al. 2013 (CRC)[29] + Ting et al. 2020 (CRC)[30] + Velez-Serrano et al. 2017 (pancreas)[31] | Surgery |
| Handling of missing features | Describe how missing features and values in the patient data sets were handled, e.g. excluded or imputed | 34 % (n = 36) | Bhandari et al. 2020 (kidney)[32] + Mourad et al. 2020 (thyroid)[33] + Smith and Mezhir 2014 (pancreas)[34] | Machine learning |
| Split or resampling | Describe how the data was split or resampled for training, validation and testing | 83 % (n = 89) | Stojadinovic et al. 2013 (CRC)[29] + Ting et al. 2020 (CRC)[30] + Velez-Serrano et al. 2017 (pancreas)[31] | Machine learning |
| Class imbalance | Make sure the data has a balanced outcome, or use mathematical measures to optimize your dataset in case of imbalance | 12 % (n = 13) | Bolourani et al. 2020 (esophagus)[35] + Bhandari et al. 2020 (kidney)[32] + Schoenberg et al. 2020 (liver)[36] | Machine learning |
| Multiple algorithms | Compare multiple algorithms to find the optimal one for a certain task | 68 % (n = 73) | Nilsaz-Dezfouli et al. 2017 (stomach)[37] + Ting et al. 2020 (CRC)[30] + Xu et al. 2020 (CRC)[38] | Machine learning |
| Multicenter | Collect data for building the algorithms from different facilities | 42 % (n = 45) | Bhandari et al. 2020 (kidney)[32] + Mourad et al. 2020 (thyroid)[33] + Stojadinovic et al. 2013 (CRC)[29] | Surgery |
| External validation set | Make use of an external validation set to assess the performance of the algorithm. This external validation set is curated from data outside the clinical environment in which the algorithm was trained | 12 % (n = 13) | Rahman et al. 2020 (esophagus)[39] + Kudo et al. 2020 (CRC)[40] + Li et al. 2020 (CRC)[41] | Surgery |
| Prospective validation | Prospectively validate decision support with new patients after training of ML algorithm was finished | 2 % (n = 2) | van Soest et al. 2017 (CRC)[42] + Adams and Papagrigoriadis 2014 (CRC)[43] | Surgery |
| Treatment recommendation | Use ML to provide a treatment recommendation. A mere prediction of complication rate is not considered a treatment recommendation | 8 % (n = 9) | Ichimasa et al. 2018 (CRC)[44] + Liu et al. 2019 (stomach)[45] + Kang et al. 2020 (pancreas)[46] | Surgery, machine learning, interaction design |
| Interface developed | Develop a user interface for clinicians and provide (online) access for other researchers | 11 % (n = 12) | Schoenberg et al. 2020 (liver)[36] + Han et al. 2020 (pancreas)[47] + Rahman et al. 2020 (esophagus)[39] | Surgery, interaction design |
| Open source | Make your code open source online for other researchers | 3 % (n = 3) | Rahman et al. 2020 (esophagus)[39] + Zhou et al. 2020 (stomach)[48] + Lei et al. 2020 (liver)[49] | Machine learning |
Study selection
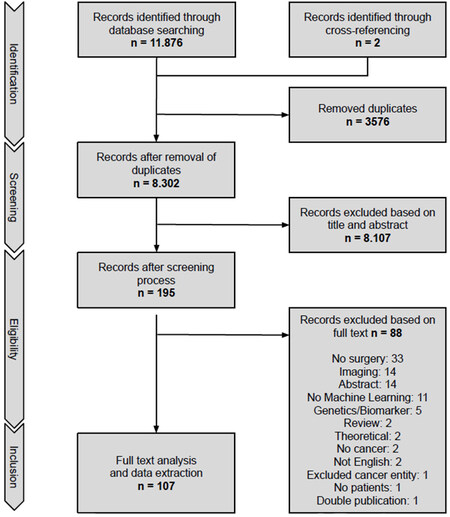
In total, 11,876 articles were identified through database searching. After the removal of duplicates and initial screening for title and abstract, 195 articles were eligible for full-text screening. In total, 107 articles met our eligibility criteria and information was extracted. The other articles were excluded for various reasons. A PRISMA flow chart of the selection process is given in Figure 2.
Study characteristics
In total, 107 studies were selected to be assessed in this systematic review; a summary for each of the selected studies can be found in Supplementary Table 2. Overall, 105 of 107 studies were conducted in a retrospective setting (98%), while only two were conducted prospectively (2%), i.e. they validated the retrospectively trained ML algorithms in a prospective cohort. No studies evaluated continuous learning of the models, i.e. investigated improvement of algorithm performance with an increasing amount of data collected during clinical use of the algorithms. Overall, 45 studies (42%) used data collected in a multicenter approach, while 23 studies (22%) used public databases such as SEER[50], ACS NSQIP[51], and SRTR[52]. External validation was performed in 13 studies (12%), with a median of 164 patients (IQR = 104-387) included in the external validation. Studies used demographic data most often (n = 98, 92%), followed by clinical data (n = 92, 86%), pathological data (n = 58, 54%), surgical data (n = 56, 52%), laboratory data (n = 55, 51%), radiological data (n = 46, 43%), biomarkers/genetics (n = 6, 6%), and radiomics (n = 2, 2%). A full list of the variables on which selection and training of ML algorithms were performed was presented by 95 studies (89%).
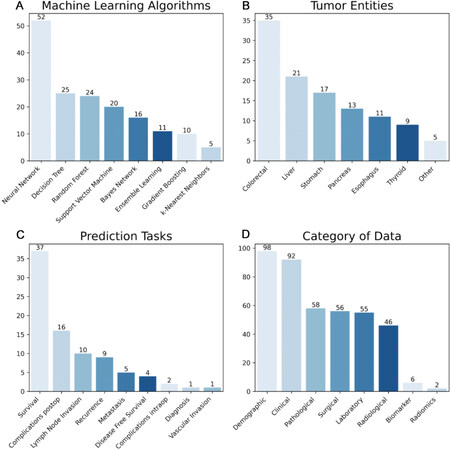
Of the selected studies, 73 (68%) used more than one ML algorithm. Neural networks (n = 52, 49%) were by far the most popular ML algorithm, only outnumbered by regression analyses (n = 56, 52%), which were used as a control method in comparison to ML methods. Neural networks (n = 45, 42%) were also the ML method most often deemed best by the authors of the respective studies. For treatment recommendations, only nine studies (8%) used ML to explore predictions that would directly influence surgical decision making. To visualize these data, Figure 3 summarizes the used ML methods, tumor entities, prediction tasks, and the origin of data.
Figure 3. Summary of study characteristics: (A) Number of machine learning algorithms investigated in selected studies; (B) number of cancer entities investigated in selected studies; (C) number of different prediction tasks performed in selected studies; and (D) origin of data in selected studies.
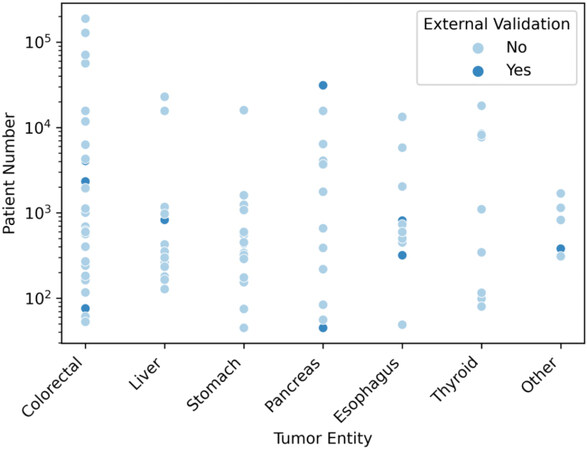
Patient numbers ranged from 45 to 188,336 (median of 565 patients, IQR = 267.5-1729.5). Figure 4 gives an overview of the different patient numbers of selected studies grouped by respective tumor entities.
Figure 4. Patient numbers of selected studies grouped by tumor entity. Tumor entities subsumed as “Other” are intrahepatic cholangiocarcinoma (n = 2), gallbladder cancer (n = 1), kidney cancer (n = 1), and peritoneal carcinomatosis (n = 1).
Methods of data splitting or resampling for validation were a single-random-split (n = 57, 53%), k-fold cross validation (n = 21, 20%), bootstrapping (n = 10, 9%), and leave-one-out-cross validation (n = 1, 1%), while 18 studies (17%) did not describe any. Only 13 studies (12%) acknowledged the problem of class balancing explicitly, and 10 (9%) of them described how they addressed it. The developed ML models were evaluated with different statistical measurements. AUC (n = 76, 71%) was the most common, followed by sensitivity
For visualization and a possible clinical application, five studies (5%) developed a nomogram and twelve studies (11%) developed an interactive interface for their chosen ML method, which was accessible online in seven cases (7%) at the time of this review. Only three studies (3%) published their code to make the methods behind their ML models accessible open source for other researchers.
DISCUSSION
AI for decision support in surgical oncology
Modern medicine generates masses of data every day, generating “big data” not only in terms of volume but also in terms of variety (e.g., demographic, clinical pathological, and surgical data), velocity (e.g., intraoperative sensor data and vital monitoring on intensive care unit), veracity (e.g., uncertainty of findings in radiological imaging), and value (for doctors, nurses, hospitals, insurance companies, etc.)[53]. However, the data themselves are useless. They have to be analyzed and interpreted, and ML has proven to be a powerful tool[54]. This systematic review gives an overview of the state of the art in use of ML and clinical application of AI for surgical oncology. Here, a broad spectrum of tumor entities investigated with various endpoints are addressed. In addition, numerous different ML algorithms are compared.
However, major drawbacks are revealed in the systematic analysis of key steps for scientific validation and clinical translation. Basic rules of machine learning were followed by most studies, such as feature set description and data splitting and resampling. Fewer studies compared multiple algorithms. Only about half of the selected studies used data from multiple centers, which is important to increase heterogeneity, reduce bias, and improve generalizability[11], as in the case of multicenter clinical trials. Moreover, up until now, prospective, external validation of ML algorithms and the development of user interfaces for clinical application of AI are limited to very few studies. The following paragraphs give an overview of how studies on AI in surgical oncology should be designed to perform an “ideal” ML study in surgical oncology based on our findings in this systematic review. These key steps lead to the path of successfully using “big data” in surgical oncology. Our checklist thus complements the DECIDE-AI guidelines that focus on early evaluation of AI systems as an intervention in live clinical settings[55]
Key steps for scientific validation
To validate ML algorithms during their development, the dataset is usually split into training and test sets. Most of the studies described their strategy for this. However, there was an inconsistent use of the terms “validation set” and “test set”, which were used as synonyms to build the algorithm. Although in many studies internal or external validations were included, they often did not describe the performance evaluation to their full extent, as reported similarly by van Soest et al.[42]. The confusing terminology and reporting lead to considerable effort in understanding the process of building surgical decision support with ML algorithms. Here, a visualization, such as a flow chart, should be used to visualize the complex process of development and validation [16]. Another weakness in the selected studies is the lack of reports on class imbalance and measures to avoid a wrongful training process. If class imbalance is not accounted for, predicting rare events or outcomes such as mortality after surgery may be distorted because, if a dataset is not equally distributed, then the ML algorithm may not predict the event but rather the majority class of “no event”.
An external validation set or independent test set is curated from data outside the clinical environment used for algorithm training and can be used to test the algorithm. External validation is a key step to demonstrate generalizability, i.e. a tool for surgical decision support cannot just be applied to the population it was designed for. However, only a few studies included in this review performed external validation.
When assessing the performance of the decision support, AUC was mostly used. Closely linked to the AUC are sensitivity, specificity, positive-predictive value, and negative predictive value. Surprisingly, results for the latter metrics are reported much less frequently, leading to considerable bias in overestimating the performance of ML algorithms in practical clinical applications of AI. A full performance analysis as well as a risk of bias analysis was not included in this systematic review as there was a lot of heterogeneity and different methods of reporting the algorithms’ performance. Instead, the data extracted on AUC - if available - for each of the selected papers can be accessed in Supplementary Table 2. The complete set of extracted performance data, including the heterogeneous metrics and methods, is available from the authors upon reasonable request. The data extracted on comprehensive performance evaluation for each selected paper is available in the Appendix. Furthermore, comparability of the studies was limited by the heterogeneity of tumor entities and predicted outcomes of the algorithms.
Multiple algorithms were compared based on about two thirds of selected papers. This approach is of utmost importance to investigate which of many different algorithms achieves the best performance for a certain prediction task and whether simpler models can achieve even better results than complicated neural networks[11,41,56].
In addition, to provide further validation after successfully developing a model, the source code should be provided online or in the supplementary material to make the surgical decision support reproducible for fellow researchers. Unfortunately, the vast majority of studies failed to provide their code.
Key steps for clinical translation
Most selected studies covered prediction purposes with no direct clinical consequence, such as predicting survival when a surgical resection was already performed. Only a few studies predicted certain outcomes that are directly linked to treatment consequences and can therefore be regarded as clinically relevant decision support. Examples include the prediction of pathological complete response after neoadjuvant chemoradiotherapy of rectal cancer that justifies the decision about a watch-and-wait-strategy[57] or the prediction of malignant intraductal papillary mucinous neoplasms of the pancreas that justifies the decision about surgical resection[46]. It has to be noted that this clinically relevant decision support renders the software used a medical product falling under the respective legal regulations.
In addition, the majority of studies described their full feature set before final feature selection. However, upon closer inspection, it was not always evident when applied features (laboratory values, results of staging, etc.) were assessed. Assessment before or after surgery influences the possible utilization for decision support: if features are only available after surgery, their application to support decisions for or against a surgical approach is limited.
Despite AI is highly acclaimed in medicine and surgery, there is a lack of tools that allow clinicians to enter patient data and then recommend individual treatment paths[8,58,59]. A famous example of that is IBM’s Watson for Oncology which gives individual advice for chemotherapy regimes for certain types of cancer. While a few studies on IBM’s Watson for Oncology have been published, the comparison with multidisciplinary tumor boards lacks the resounding success promised[59-61]. Moreover, even a seemingly strong AI can fail simply because certain chemotherapeutics are not available in the country where the system was launched[59].
Apart from that, the promising results reported in selected papers almost always lack a prospective, external validation, which is a key requirement for the transition into the clinical routine and external generalizability[35]. Often, even a randomized, controlled trial would be necessary to prove the superiority of AI-supported decisions compared to those without AI. While in this review, only a few studies on decision support in surgical oncology validated their algorithm externally, AI for imaging is two steps ahead, as these applications have not only been tested externally and prospectively but also prospectively compared with clinicians in randomized controlled trials[62].
This limitation is closely linked to the development of user interfaces that facilitate adoption into the clinical workflow[63] and allow other physicians to benefit from the findings[64]. Only when decision support is easy to comprehend as well as to operate and allows for integration into the clinical routine can it contribute to patient benefit[65,66]. On the contrary, AI and surgical decision support are not to replace physicians but to equip them with tools that may improve patient outcomes and time utilization in clinical processes. Despite popular claims that AI will solve most problems in medicine, up until now, most algorithms train models that only solve one specific problem[11].
Strengths and weaknesses
An extensive literature research of the last ten years was conducted. A large variety of different algorithms and databases was covered, yet there was still the possibility that some articles were missed. ML algorithms may not necessarily be published in clinical journals but in pre-print services, online code repositories (e.g., GitHub), or conferences[58]. It is because of the focus on the clinical translation of ML and AI that these services were not explicitly searched. Furthermore, we excluded articles investigating image analysis and only focused on surgical decisions instead of radiological decisions. This exclusion criterion may have led to the exclusion of surgical imaging methods important for surgical oncology, such as those described[67].
An in-depth bias analysis of the selected studies was not performed because of the heterogeneity of studies that rendered further clinical recommendations impossible. Nevertheless, several items, such as the description of missing features, presentation of the full feature set for data, strategy for splitting the data, class imbalance, and access to the methods online (open source), can be considered as indicators of bias.
In conclusion, this systematic review provides a detailed summary and quality checklist of AI for decision support in surgical oncology with a focus on clinical translation. There is increasing activity in this field with multiple cancer entities and different ML algorithms investigated to predict various endpoints.
Whereas most studies follow basic rules of machine learning, such as feature set description and data splitting and resampling, fewer studies compare multiple algorithms or use data from multiple centers. Moreover, until now, there are only very few studies on prospective, external validation and the development of user interfaces; however, both are necessary before clinical translation occurs and will pave the road for randomized controlled studies comparing surgeons with surgical decision support to those without such support.
DECLARATIONS
Authors’ contributionsConceptualized the study: Wagner M, Dugas M, Maier-Hein L, Speidel S, Mehrabi A, Büchler MW, Müller-Stich BP
Defined the search strategy: Probst P, Brandenburg JM, Kalkum E
Developed the methodology: Wagner M, Schulze A, Klotz R, März K, Bodenstedt S, Dugas M
Performed data curation: Schulze A, Haselbeck-Köbler M
Performed data analysis: Schulze A, Majlesara A, Ramouz A
Provided funding for the study: Büchler MW, Maier-Hein L, Mehrabi A, Müller-Stich BP
Supervised the data analysis: Probst P, Nickel F, Klotz R
Wrote the original draft: Wagner M, Schulze A, Haselbeck-Köbler M, Brandenburg JM
All authors reviewed and edited the manuscript.
Availability of data and materialsTemplate data collection forms, data extracted from included studies and data used for analyses are available from the authors upon reasonable request.
Financial support and sponsorshipThis work has been funded by the National Center for Tumor Diseases (NCT) Heidelberg, Germany within the cancer therapy program “Surgical Oncology” and by the German Federal Ministry of Health within the “Surgomics” project (grant number BMG2520DAT82), and by the German Research Foundation (DFG, Deutsche Forschungsgemeinschaft) as part of Germany’s Excellence Strategy - EXC 2050/1 - Project ID 3900696704 - Cluster of Excellence “Centre for Tactile Internet with Human-in-the-Loop” (CeTI).
Conflicts of interestAll authors declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2022.
REFERENCES
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394-424.
2. Sullivan R, Alatise OI, Anderson BO, et al. Global cancer surgery: delivering safe, affordable, and timely cancer surgery. Lancet Oncol 2015;16:1193-224.
3. Knight SR, Shaw CA, Pius R, et al. Global variation in postoperative mortality and complications after cancer surgery: a multicentre, prospective cohort study in 82 countries. The Lancet 2021;397:387-97.
4. Jones RP, Poston GJ. Resection of liver metastases in colorectal cancer in the era of expanding systemic therapy. Annu Rev Med 2017;68:183-96.
5. Keller DS, Berho M, Perez RO, Wexner SD, Chand M. The multidisciplinary management of rectal cancer. Nat Rev Gastroenterol Hepatol 2020;17:414-29.
6. Prades J, Remue E, van Hoof E, Borras JM. Is it worth reorganising cancer services on the basis of multidisciplinary teams (MDTs)? Health Policy 2015;119:464-74.
7. Hansen MFC, Storkholm JH, Hansen CP. The results of pancreatic operations after the implementation of multidisciplinary team conference (MDT): a quality improvement study. Int J Surg 2020;77:105-10.
8. Hashimoto DA, Rosman G, Rus D, Meireles OR. Artificial intelligence in surgery: promises and perils. Ann Surg 2018;268:70-6.
9. Maier-Hein L, Vedula SS, Speidel S, et al. Surgical data science for next-generation interventions. Nat Biomed Eng 2017;1:691-6.
10. Mitchell TM. Machine learning. 1st ed. USA: McGraw-Hill, Inc.; 1997.
11. Kernbach JM, Staartjes VE. Machine learning-based clinical prediction modeling - a practical guide for clinicians. arXiv 2020;11:37.
12. Willems JL, Abreu-Lima C, Arnaud P, et al. The diagnostic performance of computer programs for the interpretation of electrocardiograms. N Engl J Med 1991;325:1767-73.
13. Singh R, Kalra MK, Nitiwarangkul C, et al. Deep learning in chest radiography: detection of findings and presence of change. PLoS One 2018;13:e0204155.
14. Ahmad OF, Soares AS, Mazomenos E, et al. Artificial intelligence and computer-aided diagnosis in colonoscopy: current evidence and future directions. Lancet Gastroenterol Hepatol 2019;4:71-80.
15. Pereira SP, Oldfield L, Ney A, et al. Early detection of pancreatic cancer. The Lancet Gastroenterol & Hepatol 2020;5:698-710.
16. Christodoulou E, Ma J, Collins GS, Steyerberg EW, Verbakel JY, Van Calster B. A systematic review shows no performance benefit of machine learning over logistic regression for clinical prediction models. J Clin Epidemiol 2019;110:12-22.
17. Jin P, Ji X, Kang W, et al. Artificial intelligence in gastric cancer: a systematic review. J Cancer Res Clin Oncol 2020;146:2339-50.
18. Pantelis AG, Stravodimos GK, Lapatsanis DP. A scoping review of artificial intelligence and machine learning in bariatric and metabolic surgery: current status and future perspectives. Obes Surg 2021;31:4555-63.
19. Maier-Hein L, Eisenmann M, Sarikaya D, et al. Surgical data science - from concepts toward clinical translation. Med Image Anal 2022;76:102306.
20. Henn J, Buness A, Schmid M, Kalff JC, Matthaei H. Machine learning to guide clinical decision-making in abdominal surgery-a systematic literature review. Langenbecks Arch Surg 2022;407:51-61.
21. Garrow CR, Kowalewski KF, Li L, et al. Machine learning for surgical phase recognition: a systematic review. Ann Surg 2021;273:684-93.
22. Page MJ, McKenzie JE, Bossuyt PM, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 2021;372:n71.
23. Higgins JPT, Thomas J, Chandler J, et al. Cochrane Handbook for Systematic Reviews of Interventions version 6.2; 2021. Available from: https://training.cochrane.org/handbook [Last accessed on 22 Sep 2022].
24. Kalkum E, Klotz R, Seide S, et al. Systematic reviews in surgery-recommendations from the study center of the german society of surgery. Langenbecks Arch Surg 2021;406:1723-31.
25. Maffulli N, Rodriguez HC, Stone IW, et al. Artificial intelligence and machine learning in orthopedic surgery: a systematic review protocol. J Orthop Surg Res 2020;15:478.
27. Poole D, Mackworth A, Goebel R. Computational intelligence: a logical approach. USA: Oxford University Press, Inc.; 1997.
28. Moons KG, de Groot JA, Bouwmeester W, et al. Critical appraisal and data extraction for systematic reviews of prediction modelling studies: the CHARMS checklist. PLoS Med 2014;11:e1001744.
29. Stojadinovic A, Bilchik A, Smith D, et al. Clinical decision support and individualized prediction of survival in colon cancer: bayesian belief network model. Ann Surg Oncol 2013;20:161-74.
30. Ting W, Chang H, Chang C, Lu C. Developing a novel machine learning-based classification scheme for predicting SPCs in colorectal cancer survivors. Applied Sciences 2020;10:1355.
31. Velez-Serrano JF, Velez-Serrano D, Hernandez-Barrera V, et al. Prediction of in-hospital mortality after pancreatic resection in pancreatic cancer patients: a boosting approach via a population-based study using health administrative data. PLoS One 2017;12:e0178757.
32. Bhandari M, Nallabasannagari AR, Reddiboina M, et al. Predicting intra-operative and postoperative consequential events using machine-learning techniques in patients undergoing robot-assisted partial nephrectomy: a Vattikuti Collective Quality Initiative database study. BJU Int 2020;126:350-8.
33. Mourad M, Moubayed S, Dezube A, et al. Machine learning and feature selection applied to SEER data to reliably assess thyroid cancer prognosis. Sci Rep 2020;10:5176.
34. Smith BJ, Mezhir JJ. An interactive Bayesian model for prediction of lymph node ratio and survival in pancreatic cancer patients. J Am Med Inform Assoc 2014;21:e203-11.
35. Bolourani S, Tayebi MA, Diao L, et al. Using machine learning to predict early readmission following esophagectomy. J Thorac Cardiovasc Surg 2021;161:1926-1939.e8.
36. Schoenberg MB, Bucher JN, Koch D, et al. A novel machine learning algorithm to predict disease free survival after resection of hepatocellular carcinoma. Ann Transl Med 2020;8:434.
37. Nilsaz-Dezfouli H, Abu-Bakar MR, Arasan J, Adam MB, Pourhoseingholi MA. Improving gastric cancer outcome prediction using single time-point artificial neural network models. Cancer Inform 2017;16:1176935116686062.
38. Xu Y, Ju L, Tong J, Zhou CM, Yang JJ. Machine learning algorithms for predicting the recurrence of stage IV colorectal cancer after tumor resection. Sci Rep 2020;10:2519.
39. Rahman SA, Walker RC, Lloyd MA, et al. OCCAMS Consortium. Machine learning to predict early recurrence after oesophageal cancer surgery. Br J Surg 2020;107:1042-52.
40. Kudo SE, Ichimasa K, Villard B, et al. Artificial intelligence system to determine risk of T1 colorectal cancer metastasis to lymph node. Gastroenterology 2021;160:1075-1084.e2.
41. Li J, Gu J, Lu Y, Wang X, Si S, Xue F. Development and validation of a Super learner-based model for predicting survival in Chinese Han patients with resected colorectal cancer. Jpn J Clin Oncol 2020;50:1133-40.
42. van Soest J, Meldolesi E, van Stiphout R, et al. Prospective validation of pathologic complete response models in rectal cancer: transferability and reproducibility. Med Phys 2017;44:4961-7.
43. Adams K, Papagrigoriadis S. Creation of an effective colorectal anastomotic leak early detection tool using an artificial neural network. Int J Colorectal Dis 2014;29:437-43.
44. Ichimasa K, Kudo SE, Mori Y, et al. Artificial intelligence may help in predicting the need for additional surgery after endoscopic resection of T1 colorectal cancer. Endoscopy 2018;50:230-40.
45. Liu C, Qi L, Feng QX, Sun SW, Zhang YD, Liu XS. Performance of a machine learning-based decision model to help clinicians decide the extent of lymphadenectomy (D1 vs. D2) in gastric cancer before surgical resection. Abdom Radiol (NY) 2019;44:3019-29.
46. Kang JS, Lee C, Song W, et al. Risk prediction for malignant intraductal papillary mucinous neoplasm of the pancreas: logistic regression versus machine learning. Sci Rep 2020;10:20140.
47. Han IW, Cho K, Ryu Y, et al. Risk prediction platform for pancreatic fistula after pancreatoduodenectomy using artificial intelligence. World J Gastroenterol 2020;26:4453-64.
48. Zhou C, Wang Y, Ji MH, Tong J, Yang JJ, Xia H. Predicting peritoneal metastasis of gastric cancer patients based on machine learning. Cancer Control 2020;27:1073274820968900.
49. Lei L, Wang Y, Xue Q, Tong J, Zhou CM, Yang JJ. A comparative study of machine learning algorithms for predicting acute kidney injury after liver cancer resection. PeerJ 2020;8:e8583.
50. Jajroudi M, Baniasadi T, Kamkar L, Arbabi F, Sanei M, Ahmadzade M. Prediction of survival in thyroid cancer using data mining technique. Technol Cancer Res Treat 2014;13:353-9.
51. Merath K, Hyer JM, Mehta R, et al. Use of machine learning for prediction of patient risk of postoperative complications after liver, pancreatic, and colorectal surgery. J Gastrointest Surg 2020;24:1843-51.
52. Tanaka T, Kurosaki M, Lilly LB, Izumi N, Sherman M. Identifying candidates with favorable prognosis following liver transplantation for hepatocellular carcinoma: Data mining analysis. J Surg Oncol 2015;112:72-9.
53. Hermon R, Williams PAH. Big data in healthcare: What is it used for? Proceedings of the 3rd Australian EHealth Informatics and Security Conference; 2014.
54. Song Y, Gao S, Tan W, Qiu Z, Zhou H, Zhao Y. Multiple machine learnings revealed similar predictive accuracy for prognosis of pnets from the surveillance, epidemiology, and end result database. J Cancer 2018;9:3971-8.
55. Vasey B, Nagendran M, Campbell B, et al. DECIDE-AI expert group. Reporting guideline for the early-stage clinical evaluation of decision support systems driven by artificial intelligence: DECIDE-AI. Nat Med 2022;28:924-33.
56. Sala Elarre P, Oyaga-Iriarte E, Yu KH, et al. Use of machine-learning algorithms in intensified preoperative therapy of pancreatic cancer to predict individual risk of relapse. Cancers (Basel) 2019;11:606.
57. van Stiphout RG, Lammering G, Buijsen J, et al. Development and external validation of a predictive model for pathological complete response of rectal cancer patients including sequential PET-CT imaging. Radiother Oncol 2011;98:126-33.
60. Tian Y, Liu X, Wang Z, et al. Concordance between watson for oncology and a multidisciplinary clinical decision-making team for gastric cancer and the prognostic implications: retrospective study. J Med Internet Res 2020;22:e14122.
61. Somashekhar SP, Sepúlveda MJ, Puglielli S, et al. Watson for oncology and breast cancer treatment recommendations: agreement with an expert multidisciplinary tumor board. Ann Oncol 2018;29:418-23.
62. Nagendran M, Chen Y, Lovejoy CA, et al. Artificial intelligence versus clinicians: systematic review of design, reporting standards, and claims of deep learning studies. BMJ 2020;368:m689.
63. Kawamoto K, Houlihan CA, Balas EA, Lobach DF. Improving clinical practice using clinical decision support systems: a systematic review of trials to identify features critical to success. BMJ 2005;330:765.
64. Wu Y, Rao K, Liu J, et al. Machine learning algorithms for the prediction of central lymph node metastasis in patients with papillary thyroid cancer. Front Endocrinol (Lausanne) 2020;11:577537.
65. Moxey A, Robertson J, Newby D, Hains I, Williamson M, Pearson SA. Computerized clinical decision support for prescribing: provision does not guarantee uptake. J Am Med Inform Assoc 2010;17:25-33.
66. Bouaud J, Blaszka-Jaulerry B, Zelek L, et al. Health information technology: use it well, or don’t! AMIA Annu Symp Proc 2014;2014:315-24.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Wagner M, Schulze A, Haselbeck-Köbler M, Probst P, Brandenburg JM, Kalkum E, Majlesara A, Ramouz A, Klotz R, Nickel F, März K, Bodenstedt S, Dugas M, Maier-Hein L, Mehrabi A, Speidel S, Büchler MW, Müller-Stich BP. Artificial intelligence for decision support in surgical oncology - a systematic review. Art Int Surg 2022;2:159-72. http://dx.doi.org/10.20517/ais.2022.21
AMA Style
Wagner M, Schulze A, Haselbeck-Köbler M, Probst P, Brandenburg JM, Kalkum E, Majlesara A, Ramouz A, Klotz R, Nickel F, März K, Bodenstedt S, Dugas M, Maier-Hein L, Mehrabi A, Speidel S, Büchler MW, Müller-Stich BP. Artificial intelligence for decision support in surgical oncology - a systematic review. Artificial Intelligence Surgery. 2022; 2(3): 159-72. http://dx.doi.org/10.20517/ais.2022.21
Chicago/Turabian Style
Wagner, Martin, André Schulze, Michael Haselbeck-Köbler, Pascal Probst, Johanna M. Brandenburg, Eva Kalkum, Ali Majlesara, Ali Ramouz, Rosa Klotz, Felix Nickel, Keno März, Sebastian Bodenstedt, Martin Dugas, Lena Maier-Hein, Arianeb Mehrabi, Stefanie Speidel, Markus W. Büchler, Beat Peter Müller-Stich. 2022. "Artificial intelligence for decision support in surgical oncology - a systematic review" Artificial Intelligence Surgery. 2, no.3: 159-72. http://dx.doi.org/10.20517/ais.2022.21
ACS Style
Wagner, M.; Schulze A.; Haselbeck-Köbler M.; Probst P.; Brandenburg JM.; Kalkum E.; Majlesara A.; Ramouz A.; Klotz R.; Nickel F.; März K.; Bodenstedt S.; Dugas M.; Maier-Hein L.; Mehrabi A.; Speidel S.; Büchler MW.; Müller-Stich BP. Artificial intelligence for decision support in surgical oncology - a systematic review. Art. Int. Surg. 2022, 2, 159-72. http://dx.doi.org/10.20517/ais.2022.21
About This Article
Copyright
Data & Comments
Data
 Cite This Article 29 clicks
Cite This Article 29 clicks












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.